Antibiotics:
German scientists have discovered a new weapon to fight the hospital monster-multi-resistant staphylococcus aureus. For many years, it was hidden not in the permafrost or the Mariana Trench, but right under our noses. Or rather, in it.
In recent years, the number of infectious diseases caused by antibiotic-resistant bacteria has increased worldwide. Organisms with multidrug-resistant organisms (MDRO), such as methicillin-resistant staphylococcus aureus, enterococci insensitive to vancomycin, or gram-negative bacteria resistant to cephalosporins of the third generation, may become more frequent causes of death than cancer in the coming decades.
Antibiotics available to the population are losing their effectiveness, and their frequent and unjustified use leads to the selection of resistant forms of bacteria inside the organisms of humans and animals. MDRO in the human microbiome is difficult to detect, because usually their carriage is asymptomatic. However, under stressful conditions (after surgical operations, injuries or immunosuppression) it can develop into an aggressive infection, which will be extremely difficult to overcome. And if we also take into account the frequent resistance of such bacteria to classical disinfectants, it becomes clear why MDRO is considered a real scourge of hospitals and maternity hospitals. They are a much more tangible threat to the population of all continents than the same Ebola virus, so huge efforts are now being directed to the search and production of new antibiotics that can “crack” the protective systems of MDRO.
A few years ago, it was discovered that representatives of the human microbiota are able to produce bacteriocins that affect closely related bacteria. For example, in 2014, a new thiopeptide antibiotic, lactocillin, synthesized by ribosomes, was isolated and described from the human commensal Lactobacillus gasseri.
It may seem strange that the human microbiota produces antibiotics, because the antimicrobial search industry has always been focused on soil bacteria: it was believed that it was there that life was raging and the struggle for existence was actively going on. However, there are more than a thousand species of bacteria in the human microbiome, competing for space and nutrients. This contributes to the appearance of a real ” weapon of mass destruction — – bacteriocins produced with the help of two types of enzymes-polyketide synthases and non-ribosomal peptide synthases.
Antibiotics in the human nose
In July 2016, German researcher Alexander Zipperer and his colleagues reported that they had discovered the bacterium Staphylococcus lugdunensis IVK28 in the human nose, which suppresses the growth of methicillin-resistant Staphylococcus aureus (MRSA)
The S. lugdunensis IVK28 strain effectively fought its harmful relative only under conditions of iron deficiency and only on solid agarized media. The mechanism of the confrontation was unclear, and therefore Zipperer performed transposon mutagenesis of the cells of the isolated strain — to identify the gene responsible for the synthesis of a deadly substance for S. aureus.
As a result, it was possible to obtain a mutant IVK28, which could not suppress the growth of MRSA. Analysis of the transposon insertion site showed that it violated the structure of the gene of the supposed non-ribosomal peptide synthetase (NPS). It turned out that this gene, along with other sequences associated with the biosynthesis of antibiotics, is part of an operon with a size of 30 t.n. This indicated that the proposed inhibitor molecule may be a complex of non-ribosomal peptides.
The operon was detected by PCR in all cultures of S. lugdunensis, which means that it is characteristic of the entire species, and not only for the IVK28 strain. However, the GC composition of the operon (26.9%) differed from the GC composition of the rest of the S. lugdunensis genome (33.8%), which indicated the possible borrowing of this useful genetic cluster from other bacterial species – by horizontal transfer.
The operon consists of the lugA, B, C, and D genes encoding peptide synthetase proteins (see box below), as well as other genes whose products are necessary for the synthesis and transport of a non-ribosomal peptide.
In order to definitively impute the operon’s participation in the antibacterial activity of S. lugdunensis, the smallest gene (lugD) was removed. The ΔlugD mutant, as expected, could not suppress the growth of staphylococcus aureus, but when a plasmid with a working lugD gene was injected into it, the aggressive phenotype was restored.
Secret Weapon
The lac-operon product isolated by Zipperer turned out to be a non-ribosomal cyclic peptide consisting of five amino acids (two D-valines, L-valine, D-leucine and L-tryptophan) and a thiazolidine heterocycle. They called the antibiotic lugdunin.
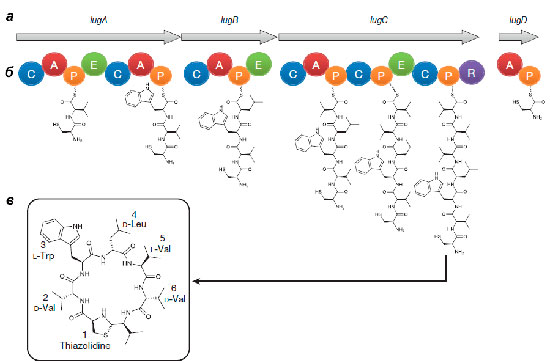
By chemical synthesis, it was possible to obtain a product with chemical properties identical to natural lugdunin and an antibacterial effect. Scientists have suggested that this antibiotic inhibits the synthesis of bacterial biopolymers-proteins, DNA and peptidoglycans.
In the experiments of German scientists, lugdunin acted not only on methicillin-resistant staphylococci, but also on glycoprotein-resistant, and even on other gram-positive bacteria such as listeria and vancomycin-resistant enterococcus. The minimum inhibitory concentration (MIC) of the new bacteriocin is 1.5-12 micrograms × ml-1, which indicates a high activity of the substance. At the same time, such concentrations did not affect the human serum in any way, did not cause neutrophil or erythrocyte lysis, and did not inhibit the metabolic activity of monocytes. Bacterial cells, however, stopped synthesizing DNA, RNA, proteins and components of the cell wall under the action of lugdunin in concentrations even lower than MIG. In this respect, lugdunin resembles daptomycin, which gives the same effect, but the mode of action of which has not yet been studied. No resistance of S. aureus cells to lugdunin was observed even after their monthly cultivation at low concentrations.
Tests in combat conditions
As expected, lugdunin’s ability to treat staphylococcal infections was demonstrated in vivo in a mouse model. In six mice, the fur on the back was shaved off and the skin was damaged by repeated gluing/peeling off the patch, Staphylococcus aureus was applied to this place. Then the skin was treated with an ointment containing 1.5 mcg of lugdunin, and six hours later the result was evaluated. Treatment with a new antibiotic greatly reduced or even completely destroyed the population of S. aureus. And not only on the surface of the skin, but also in its deeper layers.
To understand whether S. lugdunensis can prevent the colonization of the nasal cavity of vertebrates with Staphylococcus aureus in vivo, scientists conducted the following experiment. Two types of mixed cultures (S. aureus + S. lugdunensis IVK28 and S. aureus + S. lugdunensis IVK28ΔlugD) were introduced into the noses of cotton hamsters and each separately. In control cases, when one strain was injected, all three cultures stably colonized the nasal cavity. However, when introducing a mixture of S. aureus + S. lugdunensis IVK28 the amount of staphylococcus aureus after 5 days significantly decreased compared to the mixture of S. aureus + S. lugdunensis IVK28ΔlugD. This experiment showed that the production of lugdunin allows the IVK28 strain to effectively compete with Staphylococcus aureus in vivo.
It remained to figure out whether the presence of S. lugdunensis in the human nose prevents colonization by S. aureus bacteria. Zipperer and his colleagues examined smears from the nasal passages of 187 hospitalized patients. Of these, 60 people (32.1%) were found to have staphylococcus aureus and 17 people (9.1%) — S. lugdunensis. And only one patient with S. lugdunensis had S. aureus in his nose. In all isolated S. lugdunensis strains, PCR analysis demonstrated the presence of the lug operon, and all detected S. aureus strains were susceptible to lugdunin.
Lugdunin (antibiotics) was the first discovered bacteriocin of a new class. The very fact of the discovery of a new antibiotic in a representative of the human microbiota should serve as an incentive to intensify the search for other producers of bacteriocins in the composition of such communities. In the future, this will help doctors to successfully restrain the onset of multi-resistant pathogens.